Projects in our Lab
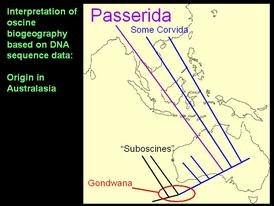 Sequence data strongly suggest the origin of all oscine passerines in Australasia, rather than just the parvorder Corvida as previously hypothesized based on DNA-DNA hybridization data.
Sequence data strongly suggest the origin of all oscine passerines in Australasia, rather than just the parvorder Corvida as previously hypothesized based on DNA-DNA hybridization data.
Passerine Phylogeny and Diversification
Higher-level phylogeny.—We are pursuing questions related to phenotypic diversification and species proliferation in passerines at multiple hierarchical levels. At the highest level, I have worked with colleagues at multiple institutions to generate a robust hypothesis of passerine relationships, at approximately the family level (e.g., Barker et al. 2002, Barker et al. 2004, Beresford et al. 2005, Barker 2014). These data have forced a major rethinking of passerine diversity patterns and diversification (e.g., Barker 2011), including completely reversing the standard scenario for the origins of the Australian passerine fauna (see figure). Ongoing work on passerines aims to expand gene and taxon sampling to produce better-resolved, temporally-calibrated passerine trees to address biogeographic and comparative questions of interest. This includes use of both next-generation and standard Sanger sequencing technologies to generate complete mitochondrial genome sequences (e.g. Powell et al. 2013) and much more extensive nuclear gene data sets for this group.
Species-level phylogenetics.—At a lower level, we are pushing forward phylogenetic studies with species-level sampling in several groups. These include the superfamily Emberizoidea (formerly known as the New World nine-primaried oscines; see tree below), a diverse group comprising ~18% of the New World avifauna, and ~8% of all birds (Lovette et al. 2010, Barker et al. 2015). This was an NSF-funded collaboration with Scott Lanyon here at the University of Minnesota, as well as Kevin Burns at UC San Diego, John Klicka at the University of Washington, and Irby Lovette at Cornell University. Other groups of current focus are the wrens (family Troglodytidae), which are an interest of long standing (Barker 2004, Mann et. al 2006, Barker 2007, Mann et al. 2009), and the mistletoe finches (Euphonia and Chlorophonia), a project of my student Tyler in collaboration with Robb Brumfield at LSU.
Higher-level phylogeny.—We are pursuing questions related to phenotypic diversification and species proliferation in passerines at multiple hierarchical levels. At the highest level, I have worked with colleagues at multiple institutions to generate a robust hypothesis of passerine relationships, at approximately the family level (e.g., Barker et al. 2002, Barker et al. 2004, Beresford et al. 2005, Barker 2014). These data have forced a major rethinking of passerine diversity patterns and diversification (e.g., Barker 2011), including completely reversing the standard scenario for the origins of the Australian passerine fauna (see figure). Ongoing work on passerines aims to expand gene and taxon sampling to produce better-resolved, temporally-calibrated passerine trees to address biogeographic and comparative questions of interest. This includes use of both next-generation and standard Sanger sequencing technologies to generate complete mitochondrial genome sequences (e.g. Powell et al. 2013) and much more extensive nuclear gene data sets for this group.
Species-level phylogenetics.—At a lower level, we are pushing forward phylogenetic studies with species-level sampling in several groups. These include the superfamily Emberizoidea (formerly known as the New World nine-primaried oscines; see tree below), a diverse group comprising ~18% of the New World avifauna, and ~8% of all birds (Lovette et al. 2010, Barker et al. 2015). This was an NSF-funded collaboration with Scott Lanyon here at the University of Minnesota, as well as Kevin Burns at UC San Diego, John Klicka at the University of Washington, and Irby Lovette at Cornell University. Other groups of current focus are the wrens (family Troglodytidae), which are an interest of long standing (Barker 2004, Mann et. al 2006, Barker 2007, Mann et al. 2009), and the mistletoe finches (Euphonia and Chlorophonia), a project of my student Tyler in collaboration with Robb Brumfield at LSU.
 Bicolored versus typical blackbirds. Birds from Black Point and Mexico exhibit a bicolored phenotype, whereas birds from Shasta Valley and Veracruz are typical of red-wings throughout most of their range.
Bicolored versus typical blackbirds. Birds from Black Point and Mexico exhibit a bicolored phenotype, whereas birds from Shasta Valley and Veracruz are typical of red-wings throughout most of their range.
Population Differentiation and Hybridization in Agelaius phoeniceus
The red-winged blackbird (Agelaius phoeniceus) is possibly the most common bird species in North America. Its ubiquity, along with its fascinating polygynous mating system has made it the focus of a massive research effort, with multiple books and untold numbers of papers dedicated to it alone. Despite this, relatively little attention has been paid to its population biology, including understanding patterns of phenotypic and genetic differentiation among populations. We are working to rectify this situation by applying modern molecular methods to large samples of these birds taken from areas of particular interest. Our work has shown that mitochondrial DNA is an extremely sensitive marker of population history in this species (Barker et al. 2012), showing strong evidence of population expansion (or possibly selection). Current work aims at differentiating selection from demographic effects using data from multiple loci. In terms of population differentiation, we recently completed a study on genetic relationships of phenotypically similar "bicolored" blackbird populations in California and Mexico (Dufort and Barker 2013; see figure). Using mitochondrial DNA and microsatellite loci developed in the lab, we have shown that bicolored populations are most closely related to one another, to the exclusion of adjacent typical populations, despite being separated by nearly 3000 kilometers.
Future work on this system will entail multilocus genetic analyses (microsatellites, sequences, and SNPs) of much more extensive population sampling from across the species range, with a focus on California and Mexico. Some basic questions include the timing of the bicolor-typical split (assuming they differentiated in allopatry), estimating the directionality of gene flow between bicolor and typical populations, and estimating local patterns of population growth across the species range. We are also interested in the influence of individual behaviors on patterns of differentiation, especially the potential effects of female choice and male-male competition related to bicolor-specific phenotypes.
The red-winged blackbird (Agelaius phoeniceus) is possibly the most common bird species in North America. Its ubiquity, along with its fascinating polygynous mating system has made it the focus of a massive research effort, with multiple books and untold numbers of papers dedicated to it alone. Despite this, relatively little attention has been paid to its population biology, including understanding patterns of phenotypic and genetic differentiation among populations. We are working to rectify this situation by applying modern molecular methods to large samples of these birds taken from areas of particular interest. Our work has shown that mitochondrial DNA is an extremely sensitive marker of population history in this species (Barker et al. 2012), showing strong evidence of population expansion (or possibly selection). Current work aims at differentiating selection from demographic effects using data from multiple loci. In terms of population differentiation, we recently completed a study on genetic relationships of phenotypically similar "bicolored" blackbird populations in California and Mexico (Dufort and Barker 2013; see figure). Using mitochondrial DNA and microsatellite loci developed in the lab, we have shown that bicolored populations are most closely related to one another, to the exclusion of adjacent typical populations, despite being separated by nearly 3000 kilometers.
Future work on this system will entail multilocus genetic analyses (microsatellites, sequences, and SNPs) of much more extensive population sampling from across the species range, with a focus on California and Mexico. Some basic questions include the timing of the bicolor-typical split (assuming they differentiated in allopatry), estimating the directionality of gene flow between bicolor and typical populations, and estimating local patterns of population growth across the species range. We are also interested in the influence of individual behaviors on patterns of differentiation, especially the potential effects of female choice and male-male competition related to bicolor-specific phenotypes.
Graduate Student Projects
Graduate students in my lab are working and have worked on a variety of projects, including:
Graduate students in my lab are working and have worked on a variety of projects, including:
- Phylogenetics of community assembly and effects of community composition on phenotypic evolution in woodpeckers (Picidae). (Student: Matt Dufort, co-advised with Ken Kozak)
- Multilocus comparative phylogeography of the birds of Baja. (Student: Hernan Vazquez-Miranda, co-advised with Bob Zink)
- Comparative analyses of cooperative breeding in birds. (Student: Michael Wells)
- Species-level phylogenetics of the blackbirds (Icteridae). (Student: Alexis Powell, advised by Scott Lanyon)
- Phenotypic evolution and diversification in New World oscine birds. (Student: Tyler Imfeld)
Undergraduate Student Projects
Undergraduate students in my lab have worked on a variety of projects, including:
Undergraduate students in my lab have worked on a variety of projects, including:
- Song variation among populations of marsh wren (Cistothorus palustris). (Student: Isaiah Clark, directed research)
- Phenotypic and genetic differentiation among subspecies of the Moriche oriole complex (Icterus cayanensis). (Student: Jacob Musser, honors thesis)
- Comparative patterns of genetic and song divergence among California populations of red-winged blackbird (Agelaius phoeniceus). (Student: Angela Pitera, directed research)
- Behavioral and genetic differentiation of Transvolcanic marsh wrens (Cistothorus palustris tolucensis). (Student: Hannah Conley, honors thesis)
- Sequencing and annotation of the Vireo olivaceus mitochondrial genome. (Student Sally Paulson, directed research)

